Webdata
Wokring with xpath and regex
After our last session where we learned how to connect R to a
database and query it, today’s session will focus on webdata and regular
expressions. First, we will use R to inspect and locate elements in html
and xml structures using xpath. Then, we will quickly look at
stringr and wrap up with regular expressions.
Both xpath and regular expressions are extremely useful when it comes to webdata. Locating data in nested html structures is particularly useful when it comes to scraping, our next session.
Regular expressions are handy whenever manipulating strings and will serve you well in many situations. In the context of webdata, they can be particularly useful when handling the type of uncleaned data we get after we scraped the internet and need to extract specific information or just tidy up our results.
HTML structure 🌳
HTML is the standard markup language for creating Web pages. It is thus important to understand the basic structure of html documents to be able to scrape particular parts of a website. HTML describes the structure of a Web page and consists of a series of elements. Elements tell the browser how to display the content, for example they label pieces of content such as “this is a heading”, “this is a paragraph”, “this is a link”, etc.
Here is an example of a document object model (DOM). Notice how the there is a cascading structure of html nodes.
<!DOCTYPE html>
<html>
<head>
<title id=1>First HTML</title>
</head>
<body>
<div>
<h1>
I am your first HTML file!
</h1>
</div>
</body>
</html>The <!DOCTYPE html> declaration defines that this
document is an HTML5 document. The <html> element is
the root element of an HTML page. The <head> element
contains meta information about the HTML page. The
<title> element specifies a title for the HTML page
(which is shown in the browser’s title bar or in the page’s tab). The
<body> element defines the document’s body, and is a
container for all the visible contents, such as headings, paragraphs,
images, hyperlinks, tables, lists, etc. The <h1>
element defines a large heading.
Developer Tools 🏄
While we can use R to inspect the parsed document, it is much easier to do this part in the browser. To do so, we right click anywhere on the website and click on “Inspect” or. On windows, you can also simply press F12.
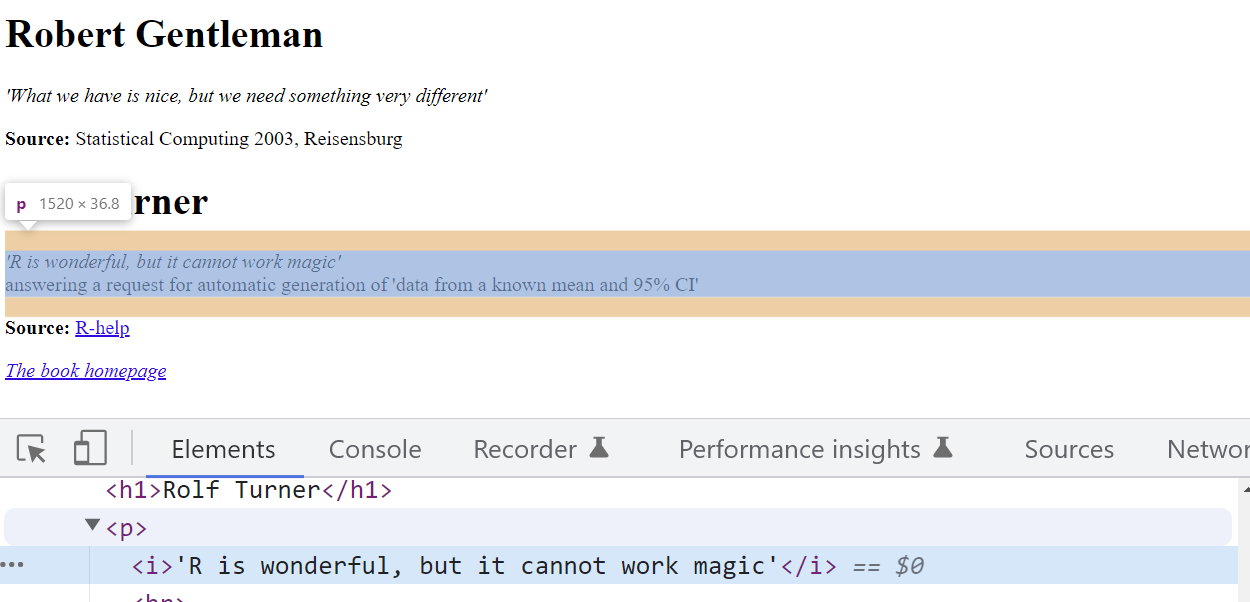
This opens up the developer tools interface on your browser. The most important tab in the developer tools is the “Elements” tab. This tab shows you the source code of the webpage in an interactive manner. Now, when we hover over the elements in the tab they will be highlighted on the webpage. By clicking the mouse icon on the top left, we can reverse this behaviour.
Parsing with R
Now let’s have a look at how to do this in R:
library(rvest)
library(stringr)You’ll learn more about rvest in next week’s session.
For now, just remember that parsing a website in R is
straightforward:
parsed_doc <- read_html("http://www.r-datacollection.com/materials/ch-4-xpath/fortunes/fortunes.html") Xpath Basics 👩💻
While HTML displays data and describes the structure of a webpage, XML stores and transfers data. XML is a standard language which can define other computer languages. XPath uses path expressions to select nodes or node-sets in an XML document. HTML and Xpath can thus be exploited in conjunction to programmtically interact with the stored HTML structure of a website.
A simple xpath in the example mentioned above would be
html/body/div/h1. The simple slashes in this example
indicate an absolute path. This means, we start at the
root node and follow the whole way down to our target element h1.
Relative paths on the other hand are indicated with
double slashes //. Relative paths skip nodes and do not
need to start at the root node. An example here would be
//body//h1.
The wildcard operator * allows us to skip elements in
the xpath.
After having parsed the webiste from HTML to an XML document, we can
locate individual elements with xpaths. The html_elements()
function from the rvest package, finds and selects elements
in the parsed document. We can use both css and xpath selectors, but for
now we will only look at xpath selectors.
html_elements(parsed_doc, xpath = "/html/body/div/p/i")## {xml_nodeset (2)}
## [1] <i>'What we have is nice, but we need something very different'</i>
## [2] <i>'R is wonderful, but it cannot work magic'</i>The xpath grammar 🧙
We can use xpath to select certain aspects of the webpage, or more precisely the underlying XML from the html file.
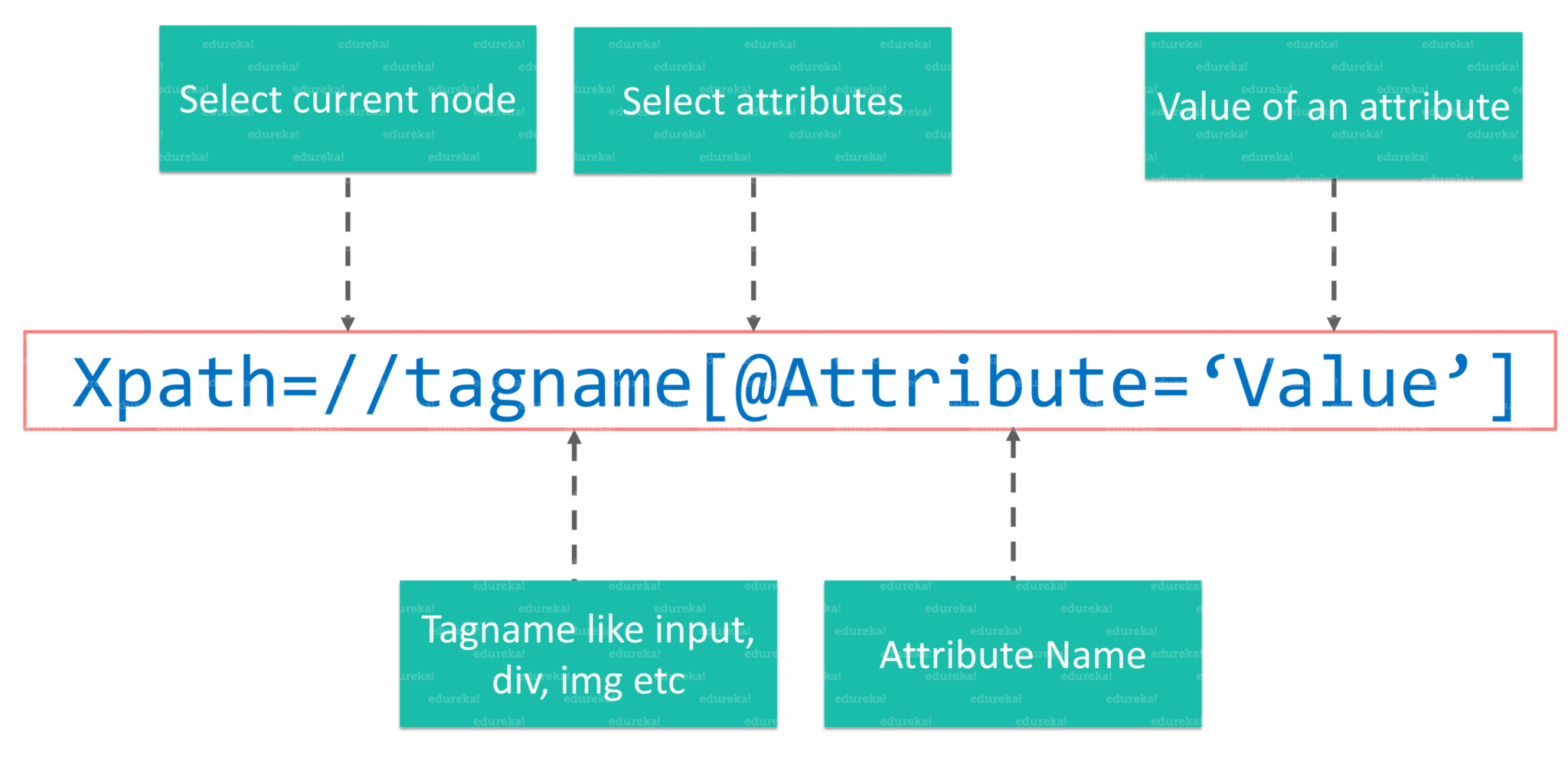

//: The releative path that lets us start with our current element- tagname: the tagname of our current element
@: The@is used to select an attribute in out element.- Attribute: The name of our attribute.
- Value: The value of our attribute
XPath Predicates
Now let’s take a look at some more complex examples of xpaths.
Elements on the webpage can also be selected with xpath by leveraging
their relations to the elements that they are connected to. The basic
syntax for this is element1/relation::element2.
If we would like to extract the two names on our example webpage, using element relations, we can do so like this:
html_elements(parsed_doc, xpath = "//p/preceding-sibling::h1")## {xml_nodeset (2)}
## [1] <h1>Robert Gentleman</h1>
## [2] <h1>Rolf Turner</h1>Finally, we can also use True/False conditions on our elements to filter them. In xpath, this is called predicates. A numeric predicate lets us select the nth element within a given path. Let us use this to extract the Source of the quotes on our example page:
html_elements(parsed_doc, xpath = "//p[2]")## {xml_nodeset (2)}
## [1] <p><b>Source: </b>Statistical Computing 2003, Reisensburg</p>
## [2] <p><b>Source: </b><a href="https://stat.ethz.ch/mailman/listinfo/r-help"> ...Next to numeric predicates, there are also textual
predicates. Textual predicates allow us to do rudimentary text
matching. This is implemented in string functions like
contains(), starts_with or
ends_with(). Predicates can also be chained together with
and. Multiple xpaths can be combined in an or logic with
the pipe operator |:
html_elements(parsed_doc, xpath = "//h1[contains(., 'Rolf')] | //h1[contains(., 'Robert')]" )## {xml_nodeset (2)}
## [1] <h1>Robert Gentleman</h1>
## [2] <h1>Rolf Turner</h1>Xpath exercise ⛏`
Can you find all links in on our example document?
Strings with stringr 💬
stringr and stringi are the two most common
libraries for string manipulation in R. Before we look into regular
expressions, let us quickly look into some of th core functionalities of
stringr and how you can use them.
Here is a quick overview of the most useful functions in the
stringr package:
x <- c("apple", "banana", "pear")
str_detect(x, "e")## [1] TRUE FALSE TRUETo extract the actual text of a match, use
str_extract(). Note that str_extract() only
extracts the first match. To get all matches, use
str_extract_all(), which returns a list.
str_extract_all(x, 'a')## [[1]]
## [1] "a"
##
## [[2]]
## [1] "a" "a" "a"
##
## [[3]]
## [1] "a"str_replace() and str_replace_all() allow
you to replace matches with new strings. The simplest way is to replace
a fixed string, however with str_replace_all() you can
perform multiple replacements by supplying a named vector:
str_replace_all(x, c("a" = "A", "b" = "B", "p" = "P"))## [1] "APPle" "BAnAnA" "PeAr"str_locate() and str_locate_all() give you
the starting and ending positions of each match. These are particularly
useful when none of the other functions does exactly what you want.
str_locate(x, 'a')## start end
## [1,] 1 1
## [2,] 2 2
## [3,] 3 3Regular expressions 📝

Regular expressions (Regex) allow us to manipulate strings based on pattern matching. Regex patterns specify a sequence of strings, either explicitly or by meta characters. While they can be a hard nut to crack, they are extremely useful.
Meta characters allow us to abstract from explicit patterns. These
meta characters are . \ | ( ) [ { ^ $ * + ?. For example,
. is called a wildcard, as it matches any character, except
for line breaks (\n).
str_view(x, "a.")Quantifiers *️⃣
? * + {n} {n,} {n,m} are so-called quantifiers,
that allow us to match the preceding character either zero or one
?, zero or more ., or one or more
+ times. {n} allows us to match a pattern
exactly n times. To match n times and more, we add a comma in the curly
brackets {n,} and to match between n and m times, we use
{n,m}.
str_view(x, "p+")We can also match for specific character classes: If you
would like to match whitespaces (spaces, linebreaks, tabs) you can use
\\w. To match any digit, simply use \\d, for
word character (digits and alphabetic characters) you can use
\\w. We can negate those character classes by using them in
uppercase (e.g. \\D matching all non-digits).
y <- "1. A small sentence. - 2. Another tiny sentence."
str_view_all(y, "\\s(\\w{4,5})\\s")str_view_all(y, "\\d\\W")R’s regex implmentation also ships with a number of predefined
character classes, like [:upper:], [:lower:]
and [:punct:]. They are usually simpler and much easier to
remember than corresponding regex patterns. Note however, that these
will require double squared brackets in many of the base R
functions.
Anchors & alternates ⚓
Next to character classes and quantifiers, anchors match the
start ^ or end $ of a string.
Alternates allow us to handle multiple cases in our pattern. As
in R |, is equivalent to an or. With [qwert]
we are creating a “one of” pattern that matches any of the strings in
the squared brackets. To negate this and get “anything but” we use
[^qwert]. Parenthesis in regex are used to create groups
and establish an order of evaluation.
str_view(x, "^a|b")fruits <- c("apple", "orange", "pear")
fruit_match <- str_c(fruits, collapse = "|")
str_subset(stringr::sentences, fruit_match) %>% head(3)## [1] "The pearl was worn in a thin silver ring."
## [2] "The fruit of a fig tree is apple-shaped."
## [3] "Canned pears lack full flavor."At this point you may wonder yourself, how we match those characters
that are used as meta characters in regex.T his is where it gets
complicated. Theoretically, we only use \ as an escape
character. However, we also use regular characters, like \d
for instance, to match digits or \s to match white spaces.
Therefore, we add another backward slash to escape the meta character
\\. To match a question mark, we would use
\\?.
z <- c('abc?defg', '123456?89', "[?.{!]\\")
str_view(z, '\\?')Finally a word of caution at the end: Since regular expressions are extremely powerful in string manipulation, it is easy to try and solve every problem with a single regex. Do not forget that you have other tools available in a programming language and you can break down the problem by writing a series of simpler regexes.
Regex exercises 🔧
- Can you explain what these regular expressions match?
"^.*$""\\{.+\\}""\\d{4}-\\d{2}-\\d{2}""\\\\{4}"
How many words are there in
stringr::wordsthat end with a “y” and are exactly 3 characers long?Now, try to find all words that end with “-ed” but not with “eed”!
Let us now write a pattern that matches both emails in the vector below.
Now try to extract all names and corresponding phone numbers from the string below.
Sources
This tutorial drew heavily on Simon Munzert’s book Automated Data Collection with R and related course materials. For the regex part, we used examples from the string manipulation section in Hadley Wickham’ s R for Data Science book.
A work by Lisa Oswald & Tom Arend
Prepared for Intro to Data Science, taught by Simon Munzert